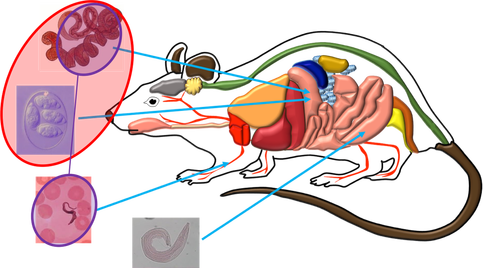
Hosts are typically co-infected by multiple parasite species, resulting in potentially overwhelming levels of complexity. A common theme of my research is the focus on within-host processes that impact parasite co-infection and host condition, with the intention of understanding how these processes feedback with processes at broader scales (population, community) to impact disease dynamics.
I propose that individual host can be considered an ecosystem, in that it is an environment containing a diversity of entities (e.g. parasitic organisms, commensal symbionts, host immune components) that interact with each other, potentially competing for space, energy, and resources, ultimately influencing the condition of the host. Tools and concepts from ecosystem ecology can be applied to better understand the dynamics and responses of within-individual host-parasite ecosystems.
I propose that individual host can be considered an ecosystem, in that it is an environment containing a diversity of entities (e.g. parasitic organisms, commensal symbionts, host immune components) that interact with each other, potentially competing for space, energy, and resources, ultimately influencing the condition of the host. Tools and concepts from ecosystem ecology can be applied to better understand the dynamics and responses of within-individual host-parasite ecosystems.
Compartmentalizing the host into discrete habitats or networks may improve our ability to predict interactions between parasites. (Rynkiewicz et al. 2015, Trends in Parasitology).
Parasite Community Assembly at Multiple Scales
Ecologists first used island systems to describe how communities of species assemble, how non-random combinations of species coexist due to competition, predation, or priority filling of niche space. Parasite-host communities are an ideal system in which to extend these ideas, with many parasites colonizing many host-islands, interacting with predators (i.e. host immune system), and limiting host resources, such as red blood cells for intracellular parasites, and colonization sites. Complexities of host-parasite systems, such as host contact rate, seasonal changes to physiology, and changes in population density, additional interesting variables for extension of established theory.
Differences in species composition and diversity between communities in similar habitats are often shaped by non-random processes. This can lead to communities of low richness being subsets of high richness communities, or nestedness. Analysis of nestedness in parasite communities has rarely been done although most hosts are co-infected. Using long-term data on wood mouse (Apodemus sylvaticus) populations in Northern England I evaluated parasite nestedness, using these results to predict order of infection in individual hosts. These results will show if population level patterns of parasite infection reflect individual experience and suggest exclusion or facilitation between parasite taxa.
Collaborators: Amy Pedersen (University of Edinburgh), Andy Fenton (University of Liverpool)
Differences in species composition and diversity between communities in similar habitats are often shaped by non-random processes. This can lead to communities of low richness being subsets of high richness communities, or nestedness. Analysis of nestedness in parasite communities has rarely been done although most hosts are co-infected. Using long-term data on wood mouse (Apodemus sylvaticus) populations in Northern England I evaluated parasite nestedness, using these results to predict order of infection in individual hosts. These results will show if population level patterns of parasite infection reflect individual experience and suggest exclusion or facilitation between parasite taxa.
Collaborators: Amy Pedersen (University of Edinburgh), Andy Fenton (University of Liverpool)
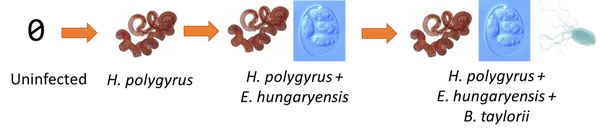
Order of infection of wood mouse parasites in individual mice predicted by nestedness analysis at the population-scale.
Wild mouse immunology in the context of parasite co-infection
|
Hosts vary in their immune response to infection, which may influence probability or severity of infection. Previous research in the laboratory and in the field has investigated the interaction between pathogen infection and immune function, but often measuring few components of the immune system. This limits ecologically relevant assessment of the complex interaction between multiple pathogens and the host immune response. I combine classic (bacterial killing assay, Rynkiewicz et al. 2013, Parasitology Research; ELISA) and novel methodologies (Luminex, transcriptomics) to study how host immunophenotype differs in relation to macro- and microparasite infection in wild mice.
The scale of the host may still be too course to quantify how parasite infection interacts with the host immune response. By measuring immune components present in specific tissues (e.g. spleen, blood, lymphnodes) we can more precisely measure how parasites impact the immune environment in different within-host habitats. This allows us to determine which parasites are more likely to interact as well as longitudinal effects on susceptibility and parasite community assembly. Collaborators: Amy Pedersen (University of Edinburgh), Simon Babayan (University of Glasgow) Using experimental infections of white-footed mice in the lab, I am using a transcriptome approach to investigate if the host immune response is mediating the facilitative interaction observed in the lab between two tick-borne pathogens, Borrelia burgdoferi and Babesia microti. The "opposite" immune profiles generated by infection with each pathogen may be the cause of the spread of B. microti following that of B. burgdorferi. Collaborators: Maria Diuk-Wasser (Columbia University) |
Microbial communities of hosts and tick vectors
|
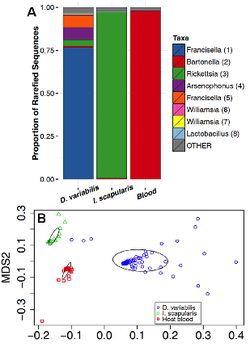
Most macro-organisms house diverse microbial communities. Of particular interest are disease vectors whose associations with microbes can have significant influences on pathogen transmission and vector competence. I have investigated the bacterial community composition and diversity of two tick species, Dermacentor variabilis and Ixodes scapularis, and blood of a shared rodent host, white-footed mouse (Peromyscus leucopus) using next generation sequencing of genomic DNA from field-collected tick and rodent blood samples to evaluate their bacterial communities (Rynkiewicz et al. 2015, Molecular Ecology).
Collaborators: Keith Clay, Clay Fuqua (Indiana University) |
Bacterial communities from two tick species (Dermacentor variabilis, Ixodes scapularis) and rodent host blood (A) were significantly different from each other (B).
|